 
Lipid identification using a MS/MS database of 120,000 tandem mass spectra (Poster) ĪSMS 2010 - 58th ASMS Conference on Mass Spectrometry Salt Lake City, Utah.Analysis of Polar Lipids in Chlamydomonas reinhardtii Using Nanoelectrospray Direct Infusion Method and Gas Chromatography and Mass Spectrometric Detection ACTA CHIMICA SINICA 2013.Journal of Chromatography A Volume 1244, 29 June 2012, Pages 139–147 Qualitative analysis of algal secretions with multiple mass spectrometric platforms.Induced Pluripotent Stem Cells Show Metabolomic Differences to Embryonic Stem Cells in Polyunsaturated Phosphatidylcholines and Primary Metabolism PLOS ONE 2012. 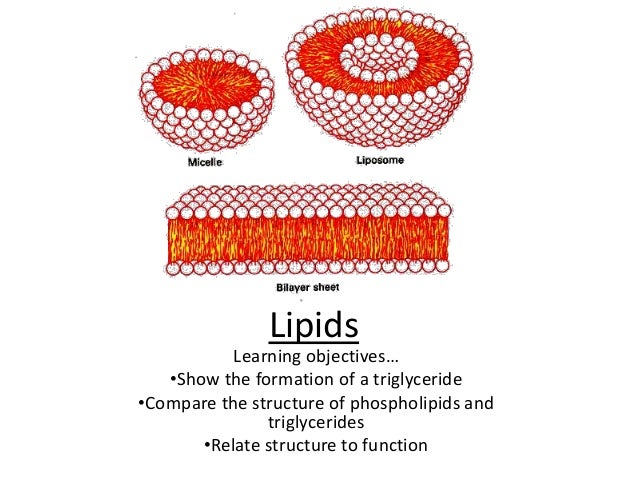
Publications and presentations using LipidBlast:ĭaily updated citation statistics from Google Scholar Download PPT and TIFF pictures (please see ZIP above). This license lets others distribute, remix, tweak, and build upon your work, even commercially, as long as they credit you for the original creation. Parts of the software supplement of the publication are published under the Creative Commons (by) license. The latest LipidBlast tools have been integrated into MS-Dial software. MS/MS-batch-search - explains how to annotate thousands of spectra in a few seconds with EXCEL output MS/MS-GUI-search - details how to search MS/MS spectra in visual mode using the NIST MS Search GUI MGF-files -explains how to create MGF files that contain all MS/MS scans and precursor ion information LipidBlast-mz-lookup - explains how to perform an MS1 search only and discusses the needed resolving power LipidBlast-Templates - this explains how to study and create lipid fragmentations using EXCEL Lipid-Classes -this link details about the covered classes and number of spectra in LipidBlast Our LipidBlast library works with low-resolution and high-resolution instruments. We developed a computer-generated (in-silico) tandem mass spectral (MS/MS) database that can be used to annotate and identify hundreds of lipids in plants, bacteria, algae, animals, humans, viruses. Lipid analysis of polar lipids can be performed with tandem mass spectrometry and mass spectral library search. PubMed Central Link (Open Access): PMC3731409 Published in Nature Methods (2013) Advanced Online LINK: doi:10.1038/nmeth.2551 Results: LipidBlast in silico tandem mass spectrometry database for lipid identification 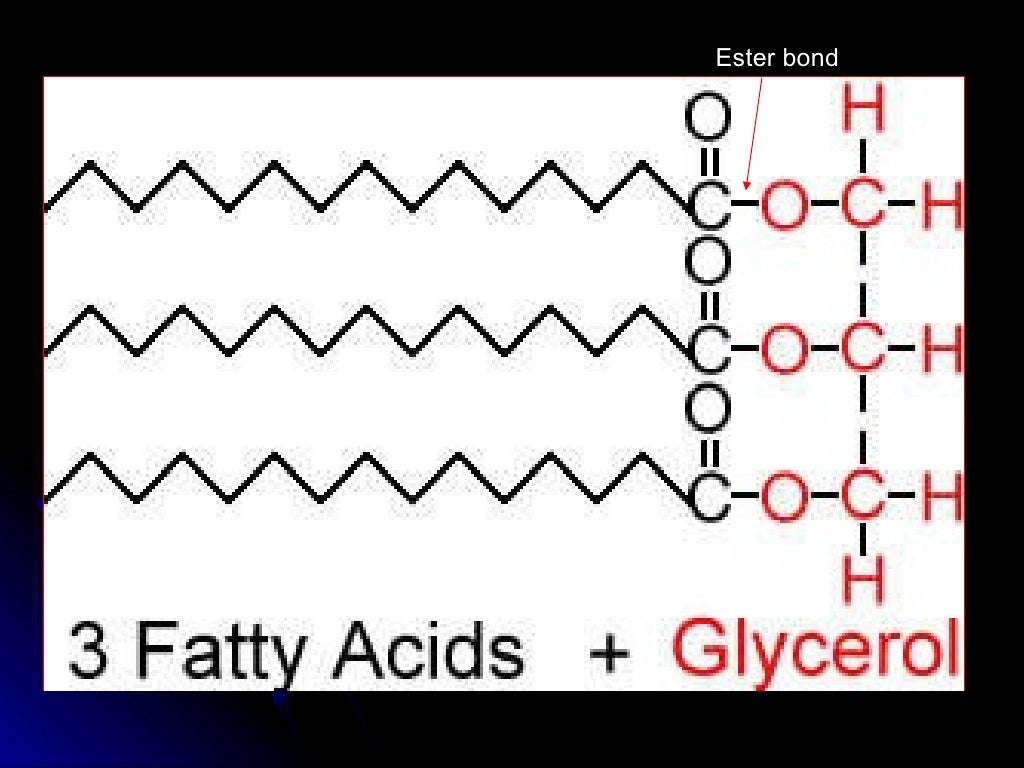
Project Partner: Tobias Kind, Kwang-Hyeon Liu, Do Yup Lee, Brian DeFelice, John K. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
